Ditching Pandas for DuckDB: The Memory Crisis Hiding in Your Validation Pipeline
It starts with a simple requirement: validate a few CSV files before they hit the warehouse. Null checks, type enforcement, range validation, basic stuff. You wire up a Python script using pandas, because that’s what the documentation shows, and it works beautifully for the first month. Then the business grows. Suddenly you’re processing 8 to 10 files daily, each containing 2 to 4 million records, and your pd.read_csv() calls start eating 16GB RAM for breakfast. At 2 AM, when the pipeline crashes with a MemoryError on a particularly chunky file, you realize the “simple” solution has become an operational liability.
This is the moment data engineering teams face the pandas-to-DuckDB migration decision. The promise is seductive: replace memory-hungry DataFrame operations with direct-to-S3 SQL queries that never touch local RAM. No cluster provisioning, no warehouse costs, just SELECT * FROM read_parquet('s3://...') and instant answers. But the migration isn’t the drop-in replacement the tutorials suggest. The reality involves network bottlenecks, Parquet decoding costs, and a critical choice between pure DuckDB and a hybrid Arrow-DuckDB architecture that can make or break your Lambda costs.
Why Pandas Falls Over at Scale
The fundamental issue isn’t that pandas is “bad”, it’s architecturally mismatched for cloud-scale validation. Pandas is single-threaded and eager-evaluation by design. When you run df = pd.read_csv(s3_path), the entire dataset streams into memory, expands to its full uncompressed size (often 5-10x the file size), and sits there while Python executes your validation logic row-by-row.
For production validation pipelines, the consensus among data engineers has shifted decisively: pandas belongs in exploratory notebooks, not in CI/CD workflows. The strict typing and vectorized execution of modern alternatives eliminate entire classes of out-of-memory errors. But the question isn’t whether to move, it’s what you’re moving to, and whether DuckDB’s S3-native querying actually solves your problem or just trades one bottleneck for another.
The Direct-to-S3 Promise (and the Hidden Bottleneck)
DuckDB’s httpfs extension offers a compelling illusion: query Parquet files directly from S3 without loading them into memory. The syntax is clean:
import duckdb
con = duckdb.connect(':memory:')
con.execute("INSTALL httpfs, LOAD httpfs;")
con.execute("""
SET s3_region='us-east-1';
SET s3_access_key_id='...';
SET s3_secret_access_key='...';
""")
result = con.execute("""
SELECT COUNT(*) as null_count
FROM read_parquet('s3://bucket/data/year=*/month=*/*.parquet', hive_partitioning=true)
WHERE user_id IS NULL
""").fetchdf()
With Hive-style partition discovery and filter pushdown, DuckDB theoretically reads only the relevant row groups, skipping irrelevant data. For small files (under 100MB), this works brilliantly. But when one engineer tested this pattern against an 807MB Parquet file representing a full year of NYC taxi data, the results exposed a harsh reality.
212 seconds
51 seconds
In benchmarks comparing execution environments, pure DuckDB took 212 seconds to process that file on AWS Glue Python Shell (1 DPU). By comparison, chDB completed the same workload in 51 seconds. The culprit? S3 scan and Parquet decoding consumed 83% of the total execution time, 176 seconds spent just pulling bytes across the wire and decompressing them, before the actual SQL query even ran.
The Arrow Dataset Intervention
Here’s the non-obvious solution that production pipelines are adopting: don’t let DuckDB handle S3 I/O at all. Instead, offload the heavy lifting to Apache Arrow Dataset, which features parallelized S3 reads, efficient Parquet decoding, and projection pushdown that operates at the row-group level.
The architecture shifts from a single-step DuckDB query to a two-stage pipeline:
import duckdb
import pyarrow.dataset as ds
import pyarrow.fs as fs
import boto3
# Stage 1: Arrow Dataset handles the brutal S3/Parquet work
session = boto3.Session()
creds = session.get_credentials().get_frozen_credentials()
s3 = fs.S3FileSystem(
region="us-east-1",
access_key=creds.access_key,
secret_key=creds.secret_key,
session_token=creds.token
)
dataset = ds.dataset(
"s3://bucket/large-file.parquet",
filesystem=s3,
format="parquet"
)
# Apply filter pushdown here, before materializing
arrow_table = dataset.to_table(
columns=["VendorID", "tpep_pickup_datetime"],
filter=ds.field("VendorID") == 1
)
# Stage 2: DuckDB handles complex SQL (JOINs, GROUP BY, WINDOW)
con = duckdb.connect(":memory:")
rel = con.from_arrow(arrow_table)
result = con.execute("""
SELECT VendorID, COUNT(*) as trip_count
FROM rel
GROUP BY VendorID
""").fetch_arrow_table()
When tested on that same 807MB file, this hybrid approach completed in 44 seconds, nearly 5x faster than pure DuckDB and even edging out chDB. The Arrow Dataset handled the S3 scan and Parquet decoding with parallel I/O, while DuckDB focused on what it does best: analytical SQL execution.
The Memory Trade-Off Nobody Talks About
But there’s a catch, and it’s a doozy for serverless environments. The dataset.to_table() call materializes the entire filtered dataset in memory as an Arrow Table. While this is faster than pandas, it can explode your memory footprint beyond the compressed Parquet size. In Lambda tests, reading that 807MB file required 8,086 MB of memory when fully materialized, dangerously close to Lambda’s 10,240 MB limit.
The mitigation is aggressive filter pushdown. By pushing predicates into the Arrow Dataset read operation, you reduce what gets materialized:
# Without pushdown: 8,086 MB memory, 15.2s
arrow_table = dataset.to_table()
# With pushdown: 3,458 MB memory, 15.2s (same speed, 57% less memory)
arrow_table = dataset.to_table(filter=ds.field("VendorID") == 1)
For truly large files, you must abandon the to_table() convenience and iterate over row groups manually, processing chunks that fit comfortably within your Lambda’s memory envelope. This adds complexity that pandas abstracts away, but it’s complexity that keeps your pipeline running.
Production Implementation Strategy
If you’re migrating a pandas validation pipeline to this architecture, here’s the battle-tested approach:
1. Replace pandas reads with Arrow Dataset + DuckDB
Don’t try to rewrite everything in SQL at once. Use DuckDB’s zero-copy Arrow integration to maintain Pythonic data manipulation while gaining SQL performance:
# Old pandas way
df = pd.read_csv("s3://bucket/file.csv")
nulls = df[df.user_id.isna()]
# New way
arrow_table = dataset.to_table()
rel = duckdb.from_arrow(arrow_table)
nulls = rel.filter("user_id IS NULL").fetch_arrow_table()
2. Leverage Hive partitioning for incremental validation
Structure your S3 paths with Hive-style partitioning (year=2024/month=03/) and enable hive_partitioning=true. This allows DuckDB to skip entire directories based on your WHERE clauses, reducing I/O costs significantly when validating date-ranged data.
3. Use MotherDuck for hybrid processing
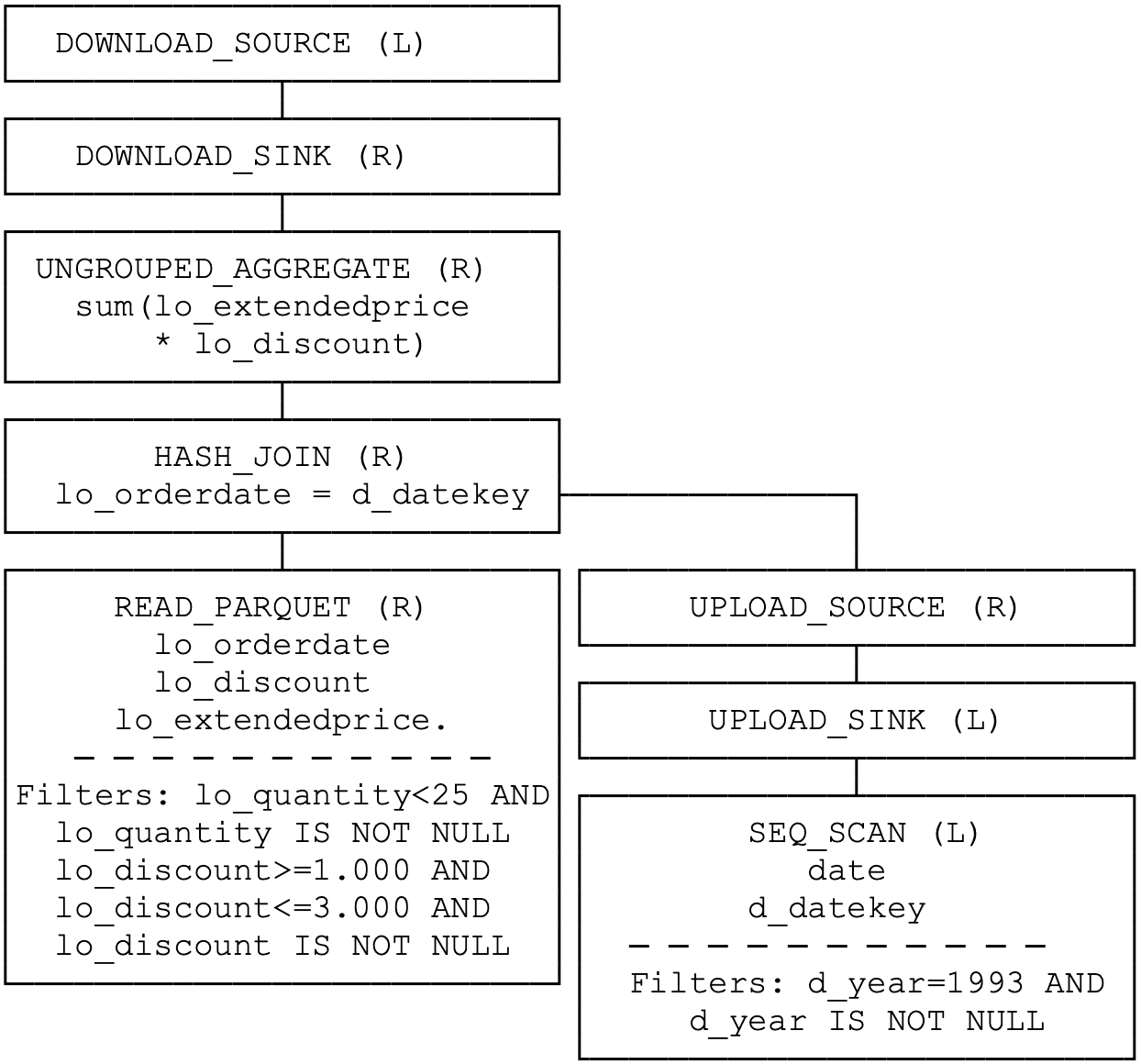
For modern data engineering pipelines that span local development and production, MotherDuck’s hybrid query processing lets you run dimension table scans locally while pushing fact table aggregations to the cloud. The “bridge operators” upload only the filtered dimension data (often megabytes) while the cloud duckling processes terabytes of fact data:
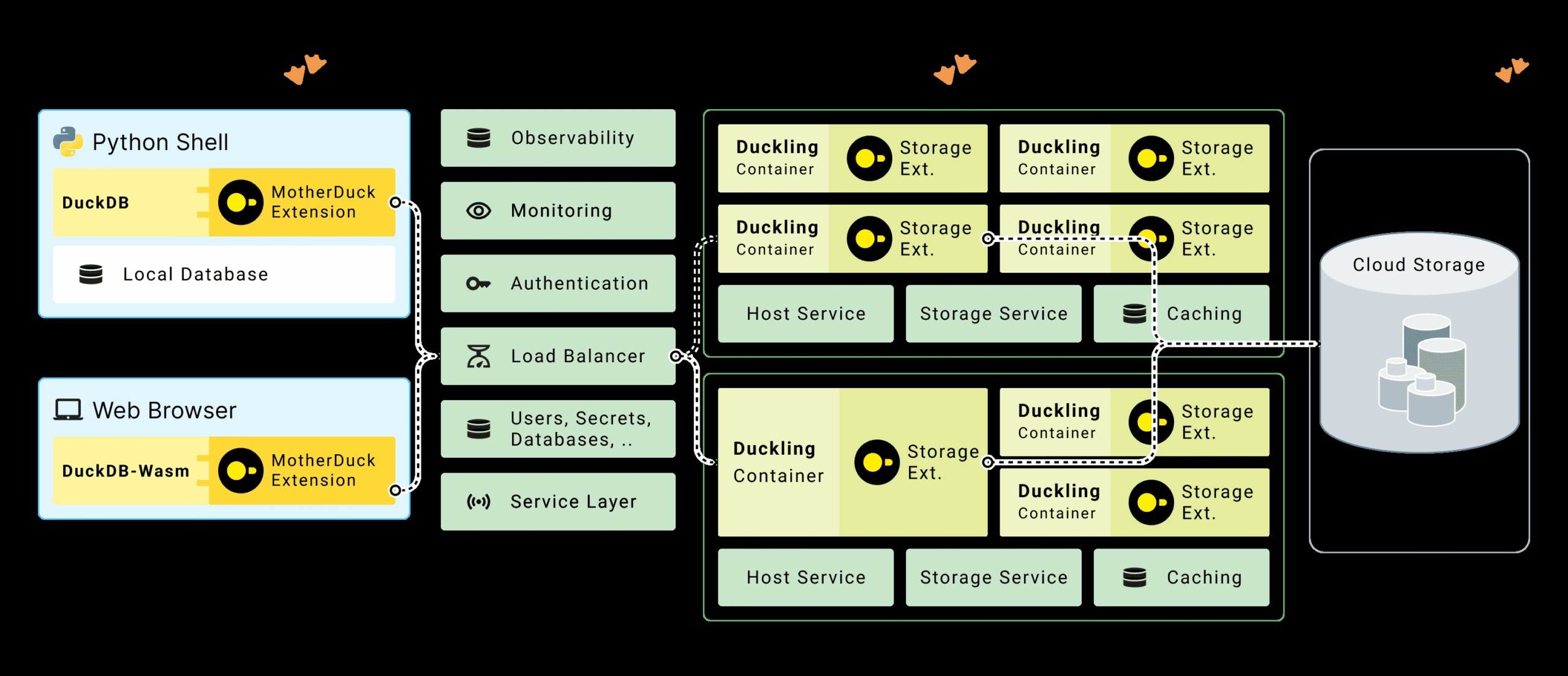
4. Memory budget for 10x file size
When sizing your Lambda or Glue workers, assume the Arrow Table will consume 8-10x the compressed Parquet size. An 807MB Parquet file required 8GB RAM. If you’re processing multi-gigabyte files, consider querying data directly from S3 without intermediate warehousing using partitioned reads, or move to Glue Python Shell with 1 DPU (which provides 16GB RAM) instead of Lambda.
When to Stay With Pandas (Yes, Really)
- Small data (< 100MB): The overhead of setting up Arrow FileSystems and DuckDB connections isn’t worth it for quick validation tasks.
- Complex string processing: Pandas’ string methods and regex operations are still more ergonomic than SQL’s
regexp_extractfor heavy text validation. - Team SQL fluency: If your data quality engineers think in Python, not SQL, forcing a language shift might cost more in productivity than you gain in performance.
For these cases, consider alternatives to traditional Pandas workloads like Polars, which offers a pandas-like API with lazy evaluation and strict typing.
The Verdict
Migrating from pandas to DuckDB for S3 validation isn’t just about swapping libraries, it’s about shifting from an “load everything” paradigm to a “pushdown and filter” architecture. The performance gains are real: 5-10x faster processing and memory footprints that scale with your query selectivity, not your total dataset size.
But the migration introduces new failure modes. The S3 scan bottleneck can make naive DuckDB implementations slower than the pandas they replace. Memory spikes from Arrow Table materialization can crash serverless functions. And the SQL translation of your validation logic requires careful testing, as SQL’s null handling and type coercion differ subtly from Python’s.
The teams winning with this migration aren’t just dropping in DuckDB, they’re architecting two-stage pipelines where Arrow Dataset handles the I/O heavy lifting and DuckDB handles the analytical heavy thinking. That’s the difference between a migration that solves your 2AM memory crises and one that just gives you different problems to debug.